New Links between Bacteria and Cancer
Microbiology and Cancer
Cancer and microbiology have been closely linked for over 100 years. Cancer patients are usually immunosuppressed due to chemotherapy, requiring special treatment and conditions to prevent bacterial infection. Bladder cancer is typically treated with inactivated tuberculosis bacteria to induce an inflammatory response which turns against remaining cancer cells, with remarkably effective results. Also, viruses are known to cause cancer, including papillomavirus (cervical cancer), Hepatitis B (liver cancer), and HTLV (human T-lymphocyte virus, causing lymphoma). In 1982, the bacterium Helicobacter pylori was discovered to be the main cause of gastric ulcers, and the first direct link between bacteria and cancer — stomach cancer — was established. The link between chronic ulcers and stomach cancer was already well known: what was not knows is that bacteria were the initial cause of stomach ulcers. Since then, several other suspects have been named, including links between Chlamydia and lung cancer, and Salmonella and gallbladder cancer.
Inflammation changes the gut ecosystem
There are two fields in which are not generally thought of as being linked: microbial ecology and cancer research. When we think of microbial ecology, we think of agricultural soil enrichment, marine ecology, air quality, nutrient recycling, species interaction, diversity and all that jazz. Not of cancer though. But in the past five years we have amassed more genomic DNA data than we have in the 50 years preceding them, including data from cancer tissue and associated bacteria. These data are beginning to show us that that the links between microbial populations and cancer are more prevalent, complex and intimate than we thought. Bacteria, as microbiologists keep repeating ad nauseam, make up 90% of the cellular population of our bodies (the extra 10% are, well, us). Following metagenomic sequencing, human microbial flora have been shown to affect conditions as varied as obesity, metabolic disease (including diabetes) infant growth and colorectal cancer — all of which we have not associated with bacteria until recently. As a result the people who study bacteria, and the people who study cancer are working together more than ever before. Last year, a study in Science led by a group from the University of North Carolina Chapel Hill, has shown a clear mechanistic link between microbial communities, inflammation, and colorectal cancer. In a nutshell, their study suggests that the following sequence of events takes place: 1) inflammation disturbs gut ecosystems; 2) this disturbance to conditions that allow pathogens to invade the gut; 3) the pathogens damage the host cells increasing the risk of the development of colorectal cancer. The study used mice that lacked the gene that makes Interleukin-10 (IL-10). IL-10 suppresses the inflammatory response, and IL10-deficient mice (IL10-/-) are genetically prone to gut inflammation. The team compared bacterial communities in the inflamed guts of IL10-/- mice with those in healthy normal (“wild type”) mice. They found that the diversity of different kinds of bacteria was significantly lower in the IL10-/- mice. But the team found little difference in microbial diversity between mice that simply had inflammation and those that had inflammation and cancer, indicating that the inflammation was the critical factor affecting bacterial populations, reducing the diversity of bacteria in the colon. In fact, one major species to shoot up and dominate the inflamed gut was E. coli, another was Enterobacter faecalis. But IL10-/- mice that were inoculated with E. faecalis only rarely developed cancer, while 80% of the group with E. coli did. Specifically, E. coli strain NC101 was foind to be the culprit. The NC101 strain has a cluster of genes under the name of “pks island”. In 2010 a group from Toulouse found that pks island genes cause cellular replication and DNA damage in the host: the harbingers of cancer. The UNC researchers colonized a mouse gut cell-line with E. coli that had the pks genes, and with E. coli lacking the pks genes. The inflammation remained, but the cells inoculated with E. coli without the pks developed fewer tumors. While mice take longer to develop tumors, the researchers saw 80% more DNA damaged cells in gut cells of IL10-/- mice inoculated with E. coli that had pks genes, versus those that were associated with E. coli without the pks genes.
Bacterial DNA in Cancer Cells
But the link between bacteria and cancer may run deeper than a changing microbial community. A recent study by a group at the University of Maryland School of Medicine shows that bacteria DNA gets transferred to human cells, in a process known as lateral gene transfer, or LGT. LGT is known to occur quite commonly between bacteria, including bacteria of different species. In fact, that is how antibiotic resistance is transferred so quickly. But findings of bacterial LGT to humans are generally treated as possible experimental artifacts, rather than true events. The UMSM team scanned data from from the 1,000 Genomes Project and found more than 7,000 instances of LGT from bacteria to human cells. When they analyzed sequences from the Cancer Genome Atlas, they discovered 691,000 more cases of LGT, and an overwhelming majority of LGT findings came from tumor samples, not from healthy cells. They found that DNA from Acinetobacter , was integrated into the genome of acute myeloid leukemia cells, especially in the mitochondrial genome. Acinetobacter is a soil bacterium but some species are known to be oppotunitic pathogens. For unknown reasons, this bacterium’s DNA was found more frequently in the genome of myeloid leukemia cells.
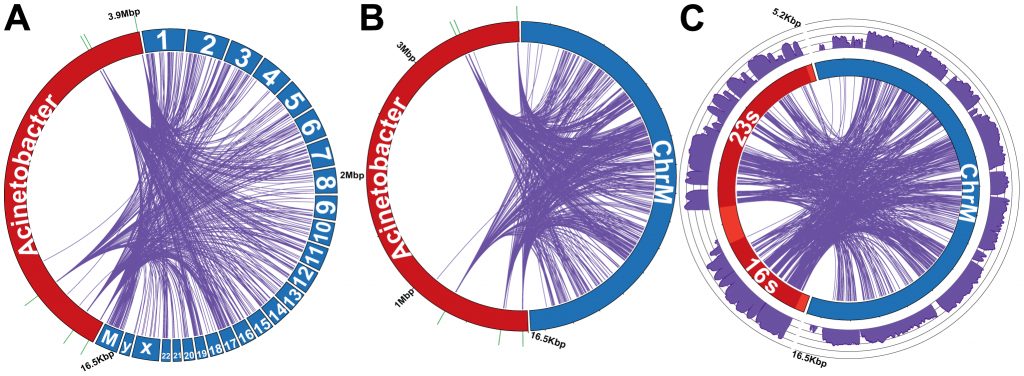
(Click for original figure) LGT from the Acinetobacter chromosome (red) to human genome (blue). Lines show which segment of DNA in Acinetobacter was transferred into which location in the human chromosome. A: all chromosomes. B: the mitochondrial chromosome, where a disproportionately large number of transfers occur. Reproduced from David R. Riley et al PLoS Computational Biology 2013
At the same time, in stomach cancer samples, the researchers found an abundance of LGT from the DNA of Pseudomonas. Here, the bacterial DNA integrated into the human DNA primarily near known oncogenes, which suggests that Pseudomonas LGT may be involved somehow in the actual progression of stomach cancer. The team also found that if the tumor had a high number of LGT’s, then the microbiome associated with it was quite uniform, comprising only a few bacterial species, primarily pseudomonas. In contrast, stomach cancer tumors with few or no LGT’s, did not have any kind of typical microbime, and the microbiomes themselves were quite diverse.
So is cancer caused by bacteria?
These are both preliminary studies. The suggested mechanism in the colorectal cancer study, and the DNA integration found in the leukemia and stomach cancer studies suggest that bacteria can potentially contribute to some types of cancer. These type of studies can be confounded by many types of laboratory based artifacts. Especially the second one: when bacterial DNA is found in human DNA data, it is typically attributed to sample contamination. The authors of the second study were very careful to address that point. (How they actually did that is worth a blog post all by itself: I found that they put large amounts of thought and effort identifying and removing artifacts, and convincing the reader of that.) Like any good science, both studies enable us to ask questions we would not have been asking before they were published. Questions such as: how does inflammation change the landscape for bacterial colonization? Can bacteria indeed harness inflammation — and then cancer — to flourish and remove competitors from their newly found ecosystem? Also, if bacterial DNA does integrate into human cancer cells, by which mechanism does that happen? Agrobacterium tumefaciens is a plant pathogen that causes tumors in plants by integrating some of its genes into the plant DNA. Could certain cancer types in humans (and other animals) be caused, or somehow exacerbated, by a similar mechanism? Finally, can we use this information to fight cancer?
Arthur, J., Perez-Chanona, E., Muhlbauer, M., Tomkovich, S., Uronis, J., Fan, T., Campbell, B., Abujamel, T., Dogan, B., Rogers, A., Rhodes, J., Stintzi, A., Simpson, K., Hansen, J., Keku, T., Fodor, A., & Jobin, C. (2012). Intestinal Inflammation Targets Cancer-Inducing Activity of the Microbiota Science, 338 (6103), 120-123 DOI: 10.1126/science.1224820
Riley, D., Sieber, K., Robinson, K., White, J., Ganesan, A., Nourbakhsh, S., & Dunning Hotopp, J. (2013). Bacteria-Human Somatic Cell Lateral Gene Transfer Is Enriched in Cancer Samples PLoS Computational Biology, 9 (6) DOI: 10.1371/journal.pcbi.1003107



















There also the use of Listeria in metastatic lung cancer: http://www.pnas.org/content/110/21/8668.abstract and many others.
I wonder what the implications are for soil health and cancer prevention? We are what we eat, right? Therefore, aren’t we what we eat eats also?
[…] * A new study finds more links between bacteria and cancer. […]
[…] A recent study by a group at the University of Maryland School of Medicine shows that bacterial DNA gets transferred to human cells, in a process known as lateral gene transfer, or LGT. LGT is known to occur quite commonly between […]
[…] recent study by a group at the University of Maryland School of Medicine shows that bacterial DNA gets transferred to human cells, in a process known as lateral gene transfer, or LGT. LGT is known to occur quite commonly between […]
[…] A recent study by a group at the University of Maryland School of Medicine shows that bacterial DNA gets transferred to human cells, in a process known as lateral gene transfer, or LGT. LGT is known to occur quite commonly between […]
[…] Read More: bytesizebio […]