So what’s new with humans?
Man is the only animal that laughs and weeps, for he is the only animal that is struck with the difference between what things are and what they ought to be.
— William Hazlitt
We like to think that we are the only species capable of emotional self-awareness and therefore the only “animal that laughs and weeps”, but that is quite probably untrue, as other animals have been shown to laugh and perhaps weep.
Whatever that elusive quality is that distinguishes us from our closest cousins, the chimps and the bonobos, it is to be found in our genome. Since human and some great apes and other primate genomes have been sequenced, the basis for comparing these blueprints exists. Many studies have been done comparing the conservation of genes, copy numbers of genes, intergenic regions, control regions, synteny, splicing and other mechanisms that may explain the differences between us and our 96% cousins. As expected, no one factor can explain why bonobos are peaceful and sexual, chimps are aggressive and patriarchal, and humans worry about taxes and blog.
Are there any new genes in humans that can help explain these differences? New genes can arise in various ways: gene duplication, exon shuffling, horizontal transfer, genes may split up (fission) or merge (fusion).
But how about genes that are completely new in humans? Do we have genes that we can claim as our own and are neither homologous to those in other apes nor have arisen from a mix & match manipulation in the common lineage of all apes? Are there actually human genes that are just that: exclusively human?
A group from China and Canada has decided to tackle that question. They looked specifically for genes that are new in the human lineage, but not in chimp or orangutan. (I’m not exactly sure why they did not look in Gorilla too, which is the other great ape with a mostly sequenced genome, perhaps because the assembly is still very much in progress.)
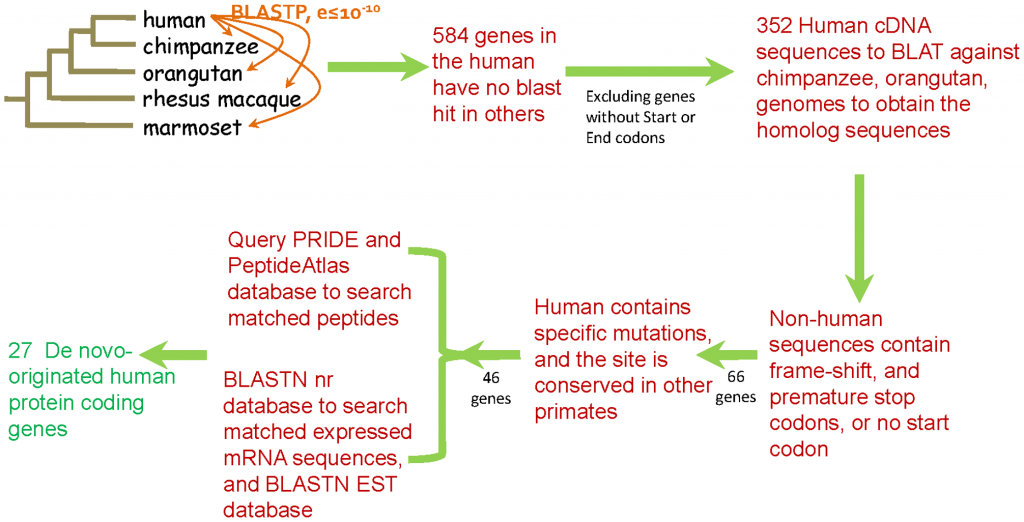
So how does one go about looking for genes that are human-only? The pipeline Wu and colleagues have set up looks like this:
Clockwise, from top left:
1. They scanned the human genome for genes with a high similarity in the genomes of chimp, orangutan and rhesus macaque. That left them with 584 genes (out of roughly 25,000) which did not have an ortholog in other primates.
2. A simple sanity check: those human genes with no start or stop codons were probably mis-identified. We are now down to 352 genes.
3. Of the 352, they looked for those that have disrupted homologous regions in chimp and/or orangutan. That mans that while the gene is functional in humans, it is not functional in the other primates. Disrupted homologous regions can mean that in non-humans the gene does not have a start codon, or has a premature stop codon, or has some frameshift mutation that renders it non-translatable. From 352 we are now down to 66 new human gene candidates.
4. But a human gene, even if not functional in other primates, may have been functional in a common ancestor of all primates, lost in the orangutan and chimp lineages, but maintained in humans. This history not make the gene as brand-new human-only. So in the 66 remaining genes they looked for sequences where the mutation that rendered them functional (like an ATG start codon, or the removal of a missense mutation) was found only in humans. Now we are left with 46 genes.
5. Great, so we have 46 open reading frames in humans that look like original, human-lineage only genes. But are they functional? Do they actually transcribe into RNA and translate into protein? (RNA-only genes were excluded from this rather conservative pipeline, they are hard enough to identify as it is.) To find that out, they looked for transcribed regions EST databases (for RNA), and in the PRIDE peptide database (for protein). Now we are left with 27 genes that are novel in humans, and because they are translated are probably active.
Trouble is, some of these genes are listed only in certain versions of Ensembl, the genome database from which the researchers took their data; (they used version 56.) This highlights a problem with the annotation of genes with no homologs: their annotation is volatile, and may change between different versions of the same database of the exact same genome. To overcome this problem, the researchers subjected different versions of Ensembl (40 through 55) to the same pipeline described above. They discovered an additional 33 genes that are candidates for de novo human-lineage only active genes, bringing the total up to 60.
What are those genes like? Why are they found only in humans? Can they help explain the differences between human and other primates? Well, for one, they’re short. Only one or, at most, two exons. This makes sense as these relatively new genes had not the time to accumulate splice sites.
The researchers moved on to look where the genes were expressed. They used RNA-Seq data from 11 different human tissues: adipose, whole brain, cerebral cortex, breast, colon, heart, liver, lymph node, skeletal muscle, lung and testes.
Here is what they found:
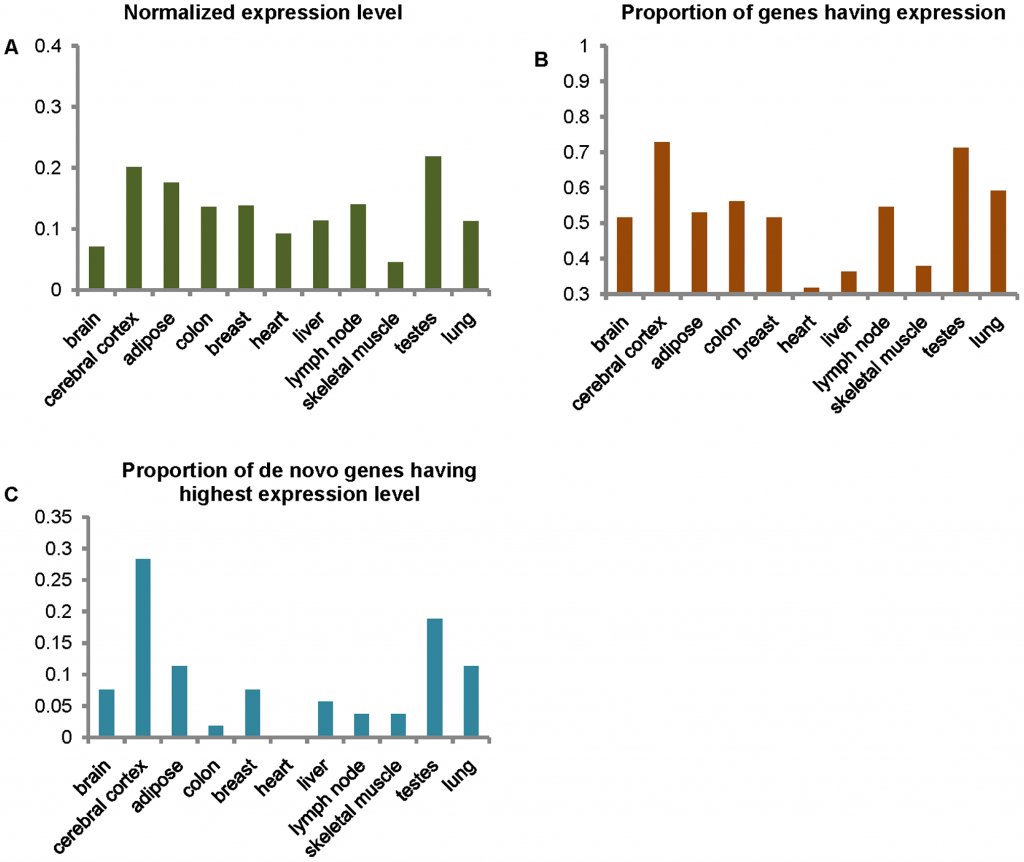
- Levels of expression of de novo genes in 11 tissues. (A) Mean normalized expression levels of de novo originated genes in 11 tissues are defined by the mean level of expression as the numbers of unique reads mapping to coding regions divided by the total length of all the coding regions, divided by the total number of valid reads in the samples (×10−8). The vertical axis represents value of mean the normalized expression levels and abscissa axis represents the 11 tissues. (B) The proportion of the de novo originated genes that have expressed reads in the 11 tissues. The vertical axis represents the values of proportion, and abscissa axis represents the 11 tissues. (C) The proportion of the de novo originated genes having their highest normalized expression levels in each of the 11 tissues. The vertical axis represents the values of proportion, and abscissa axis represents the 11 tissues. doi:10.1371/journal.pgen.1002379.g002
Panel C is the business bit: the expression of the 60 de novo human genes normalized by the general expression levels of genes in those tissues. (Pray, where are the error bars?). Seems like in Woody Allen’s two favorite organs, the testes and the cerebral cortex, do these genes have the highest expression. This actually makes some sort of sense: the testes are hypothesized to be a hotbed (sorry…) of evolutionary novelty, with all the meiosis going on there. The high expression of the de-novo human genes in the cerebral cortex also seems to confirm our anthropomorphic prejudice: we are smarter. Yay. EDIT: Following MRR’s comment: yes, we should check de-novo genes and their expression in chimps. Perhaps the high expression of de-novo genes exclusive to chimp lineage is in the cerebral cortex and testes too.
The authors do point out that there may be many other de-novo human lineage genes:
Our estimated rate, though, for de novo origin may be underestimated due to the conservativeness of our pipeline. First, as described above, in our pipeline, translatable open reading frames must have been complete in the human genome and disrupted in both the chimpanzee and orangutan genomes to be candidates as a de novo gene. Genes that did not have a clear ortholog (i.e., a sequence with very high similarity) in either the chimpanzee or the orangutan genomes (both of which are less complete than the human genome, and thus could be a missing genes) were not used. It is also often difficult to determine whether a protein-coding gene originated specifically on the human lineage or if it originated in a primate ancestor but was then lost on both the chimpanzee and orangutan lineages. The conservativeness of our pipeline thus only allowed us to accept genes where we could clearly show human specific mutations generated complete protein-coding reading frames, and that these were conserved for disrupting state in both the chimpanzee and orangutan genomes. As both the chimpanzee and orangutan sequences should be non-functional sequences, and thus not under selection, there is a reasonable likelihood that a second mutation, in addition to the human open reading frame completing mutation, could have occurred in the chimpanzee or orangutan that would prevent us for identifying these genes as having a de novo origin on the human lineage.
Also, PRIDE and PeptideAtlas, the databases of proteins they used may be underpopulated, and not include many other proteins.
To conclude, yes, humans do have their own brand-new genes which, together with many other genomic features, may help explain the differences between humans and other primates. And there are probably more of these genes than we have found so far.
As for what it means to be human:
Far out in the uncharted backwaters of the unfashionable end of the Western Spiral arm of the Galaxy lies a small unregarded yellow sun. Orbiting this at a distance of roughly ninety-eight million miles is an utterly insignificant little blue-green planet whose ape-descended life forms are so amazingly primitive that they still think digital watches are a pretty neat idea.
Perhaps it was the late, great Douglas Adams who nailed it.
Wu, D., Irwin, D., & Zhang, Y. (2011). De Novo Origin of Human Protein-Coding Genes PLoS Genetics, 7 (11) DOI: 10.1371/journal.pgen.1002379




















So what’s new with Chimps?
I feel that such studies usually miss symmetry by not investigating with the same methodology new genes (or whatever other novelty they are into to) in other species, notably chimps and gorillas. Do we have more new genes which are expressed in the brain, or is it a general trend of novel genes in mammals (in vertebrates?) to be expressed in the brain? As Henrik’s work you linked to shows, it certainly is a general trend for testis, nothing human-specific.
Of course the difficulty is that the other genomes are much less well assembled than the human genome, and dependent on the human genome for annotation, so finding novelties there will be harder.
Still, I regret the continuing human-centricity of this research program.
Hey there, cool post – have you thought about entering anything into the monthly molecular biology blogging carnival? It’s being hosted over at my blog (Rule of 6ix) next month if you’d like to put something up. Check the link here: http://blogcarnival.com/bc/submit_10473.html
Connor
@MRR there’s also the lack of EST, RNA-Seq and peptide data in apes. But running the genome-only part of the pipeline on chimps should be interesting.
On another matter, they probably need to look at more human genomes to ascertain that the genes they found only have one copy. Copy number variation exist for quite a few genes. Even between Venter’s and Watson’s genomes.
@Iddo yes and there’s also copy number variation in chimps and gorillas:
http://genome.cshlp.org/content/21/10/1626.long
While there may be human genes that are not expressed in our primate cousins, this does not mean that they are “exclusively human”. They may well be present in our more distant ancestors. There are certainly many genes that are defective or missing in humans and present in the other great apes and other mammals, including, for example, those that act in the synthesis of vitamin C. In fact, some of these genes like myosin and Fox P2 genes, due to their defects, may have actually facilitated the development of larger brain size and speech. In general, the brain and testes often show the most diverse patterns of gene expression. This is particularly evident from the analysis of gene expression available on the open-access TranscriptoNET (www.transcriptonet.ca) website produced by Kinexus. Consequently, it is not surprising that the 60 de novo gene identified showed up more frequently in these organs. The concept that the high expression of some of these genes may account for our “greater” intelligence is also questionable. The definition of intelligence is very subjective. The elephant brain is about 3-times larger than humans. We know from human developmental studies that if neurons are not innervated during early development after birth, they undergo apoptosis. Neurons that are not used become lost. It would seem that elephants, and perhaps even more so whales, must be doing a lot more with their brains than we give them credit for.
Great post, and very well written. Thanks so much for sharing!
@S.Pelech-Kinexus
While there may be human genes that are not expressed in our primate cousins, this does not mean that they are “exclusively human”. They may well be present in our more distant ancestors
The authors addressed that by looking for mutations that appear exclusively in the human lineage. (Step #4 in the pipeline.)
They may well be present in our more distant ancestors. There are certainly many genes that are defective or missing in humans and present in the other great apes and other mammals, including, for example, those that act in the synthesis of vitamin C
The ascorbate synthesis pathway is not present, as far as I know, in any of the great apes. It appears to have been lost multiple times in mammals, as bats and guinea-pigs also lack the pathway (but many other mammals do have it).
In general, the brain and testes often show the most diverse patterns of gene expression. This is particularly evident from the analysis of gene expression available on…
Yes, but as MRR pointed out, we need quantitative tissue-specific transcription data from other primates, not just human. Hopefully we will have that soon
In their publication, the authors did not apparently try to search for the existence of their 60 de novo human genes in any non-primates. I blasted the amino acid sequences of a couple of these genes against the UniProt databases and actually observed some decent matches in other species. It would seem that some of these genes may have recovered functionality in humans.
Humans lack a functional gene for the last step of vitamin C synthesis. A similar situation also exists in guinea pigs. I am not surprised that as more species are examined in this respect, this may be more common occurrence.
Of the approximately 200 different cell types in humans, about half of these are believed to exist in the brain. This probably accounts for the higher variation in gene expression patterns observed in this organ.