Reading entrails, 21st-century style
And the future is certain Give us time to work it out -- Talking Heads - Road to Nowhere
We are a species obsessed with knowing what the future holds. Our personal future, the future of our kith and kin, our countries, and our planet.
Humans have always been trying to predict their personal future. Palms, stars, cards, dreams, knuckle-bones, coffee grounds, tea leaves, bird flight patterns, crystal balls and animal entrails have all been used (and many are still in use) for predicting the future. As we consider ourselves (industrialized nations) to have matured somewhat beyond this pish-posh, we have adopted so-called scientific methods for predicting future events. Even palmistry sounds better when advertised by Professor Mirza!
A common belief held by society is that parts of our personal future are, to borrow a Calvinistic term, predestined. Personal determinism is an underlying assumption that we encounter when we go looking for a job or enroll to a school. Psychometric tests and IQ tests claim to predict our ability to succeed at post secondary education. Aptitude and personality tests claim to predict our ability to perform at a certain job, how aggressive, innovative, timid, social or deviant we are. If we know the future, we can be prepared to deal with it, we may even be able to change it. From Oedipus Rex to Terminator III, human culture is imbibed with stories of those who, armed with the gift of prescience have changed a future and those who failed miserably, for better or worse. We are creatures of certain aptitudes, talents, and as yet undeveloped skills. The deterministic view has it that we can only hope to find out what they are, and develop them to the best of our ability. There may be some wriggle room within our assigned compartment, but it is still a compartment, and it is still assigned.
The future ain’t what it used to be
Recently, genomics emerged as a mechanism for a personal future prediction, or rather unveiling our ultimate personal determinism, but also giving us the ability to forestall or change certain outcomes that are supposedly branded in our genome. Since the advent of the human genome project, the popular media has been constantly bombarding us with such metaphors as: “the blueprint of life” and “unlocking our lives’ secrets”. We expect our genome to be read like the tapestry of life woven by the Norns: a document that, when properly read, can tell us our future. Which illnesses shall befall us, and when can we expect them? Would our unborn children be healthy? Intelligent? Musical? Delinquent?
Consequently, the scientific entrails-reading business is booming. deCODE, 23andme, Bioresolve, Navigenics and other personal genomics companies offer to analyze common variants of their customers’ genome. But the personal genomics most of these companies offered is not “genomics” in the strict sense of the word. They do not analyze the full genome of their consumer. The return-postage paid cheek swab gets analyzed for common population disease markers, which cost $400-$1500 US, rather than sequencing the whole genome, which is considerably more expensive, and as we shall see, cannot at this time, provide us with much more information than analysis of common variants.
Therefore for now, the personal future prediction is limited to prediction of susceptibility to diseases as revealed by common variants in the population. In few cases the prediction is unequivocal: a G6PD deficient individual should should not eat fava beans; a person with more that 40 CAG repeats in the Huntingtin gene will develop Huntington’s disease. Most predictions are probabilistic. Someone with 36-39 CAG repeats in Huntingtin may or may not develop the disease. Women with certain BRCA1 mutations have an 85% probability of developing breast cancer and a 55% probability of ovarian cancer by the time they are 70. Several mutations in the TCSF7L2 gene are associated with type 2 diabetes.
But most genetic diseases or phenotypes are not monogenic. The genomic predictors are typically spread between several loci, or many loci, or, as we have come to realize lately, a very large number of loci. Genome wide association studies have revealed that, if anything, only a few diseases are explained by looking at common population variants. Even then, the combination of common variants usually predicts only a low single digit increase in susceptibility.
The New England Journal of Medicine published a series of opinion pieces addressing these findings, and what they mean for the future (“future”, again!) of human genetic disease studies. Dr. David Goldstein is the director of the Center for Human Genome Variation, Institute for Genome Sciences and Policy at Duke University. In his opinion, genome wide association studies of common genomic variants have pretty much run their course. Although highly successful in a few cases, for most cases they fail to explain the genetic components of diseases. His conclusion is that it is time to look at a much larger number of variants, which include rare population variants. This would mean sequencing whole genomes of selected individuals. Peter Kraft and David Hunter differ in their opinion, and claim that there is still much to be discovered in the study of common variants, and we still have quite a way to go, not only in discovering more common varaints, but also in tuning the statistics of the tests we already have. Joel Hirschhorn warns that disease risk assessment is only a by-product of genome-wide association studies, and that the chief goal should be understanding the biological pathway components that are part of these diseases.
Prediction is very difficult, especially of the future.
The cost of DNA sequencing is dropping exponentially, which means that true personal genomics — affordable whole genome sequencing — will very soon be here. Prof. Steven Brenner from University of California Berkeley is already pre-empting the move in personal genomics from common variant analyses to whole genome analyses. In Prof. Brenner’s opinion, we are woefully unprepared for the deluge of genomic data that will accompany true personal genomics. Even today, when we have the full sequences two human genomes publicly available, the low informational return from these data has more to do with data mismanagement than anything else.
The effects of mutations are scattered amongst hundreds of databases and
amidst millions of manuscripts and patent applications. These data are
heterogeneous; while some papers discuss the precise effects of a
single nucleotide change, many analyses basically offer rules of thumb.
All such information could be powerful in personal genome
interpretation, if only we could make use of it.
-- Steven E. Brenner, http://www.genomecommons.org
Brenner suggests a managed crowdsourcing approach to curating all this knowledge into a single repository, the Genome Commons. The human genome will be annotated and curated by a large group of experts. At the same time, even if that information would be properly curated, once we go beyond common markers to whole genome analysis our chief problem would be avoiding being drowned by “genomic marginalia”. Some of these issues,particularly those of genomic input are being addressed by Human Variome Project, of which I have written in an earlier post.
Not a book
Regardless of one’s view on personal determinism, it seems that the “Book of Life” analogy is a misleading one. We are not even reading all the letters: methylated cytosine, the “fifth base” still eludes most sequencing technologies, and we know that methylation is a factor in the expression of genes. Alternative splicing is still a rather opaque topic. Even if at some future date we could read our genome, by amassing a huge effort to make it legible, and correlating many variants with diseases and other phenotyopes, we would only enlarge a set of fuzzy pictures of probable futures. Our genomes are not a books to be read linearly. Any genome is a set of interacting set of intricate and non-deterministic probability functions. Moreover, the nature of these functions change over time based on how we live: what we eat, breath, drink, do, and with whom. Nevertheless, we are an inquisitive species and we strive to be in control of our individual and collective futures. We would keep collecting sequence data and refine our understanding of how newly discovered probability functions may affect our lives. The hazards of one sided interpretation of findings are well known to any scientist, detective or physician that are worth their salt. While we are in the process of amassing and ordering the human genomic data in years and decades to come, we should put an equal effort into the less palpable goal of learning how to interpret and, more importantly, how not to over-interpret our findings.
Brenner, SE. (2007). Common sense for our genomes Nature, 449 (7164), 783-784 DOI: 10.1038/449783a
Hardy, J., & Singleton, A. (2009). Genomewide Association Studies and Human Disease New England Journal of Medicine DOI: 10.1056/NEJMra0808700
Goldstein, D. (2009). Common Genetic Variation and Human Traits New England Journal of Medicine DOI: 10.1056/NEJMp0806284
Hirschhorn, J. (2009). Genomewide Association Studies — Illuminating Biologic Pathways New England Journal of Medicine DOI: 10.1056/NEJMp0808934
Kraft, P., & Hunter, D. (2009). Genetic Risk Prediction — Are We There Yet? New England Journal of Medicine DOI: 10.1056/NEJMp0810107
Nicholas Wade (2009). Genes Show Limited Value in Predicting Diseases New York Times






















This is a very interesting perspective! I think part of what draws people to sci-fi are the ethical dilemmas posed when a society’s scientific capabilities exceeds what the society is ready for — in this case, our genetic tools exceeding our current ability to treat/quantify (or even the common man’s ability to understand) the risks that our new capabilities can assess.
What’s the solution? I’m not sure there’s an easy one, but at the most basic level we should invest in better genetics education in high school so that people don’t succumb to fallacious claims about how they have the “gay gene” or some other equal nonsense…
There’s a lot of “reading the entrails” that goes on, but it’s easy to isolate those companies. 23andme and deCODE aren’t the entrail reading kind of companies. They provide you with your information, and whatever interpretive comments are supported by the scientific literature.
An important point to me is that even if no interpretation were being done, the service is still valuable. Full genome sequencing would certainly be better(and there are companies working on this), but I’ll take what I can get and along with that I’ll take responsibility for not over-interpreting. At least I have the option, instead of having to ask a doctor for permission to find out about myself(caveats about the quality of the info withstanding).
Proteomics might be coming to the consumer if this genomics stuff works out, so I’m really excited for what the future holds.
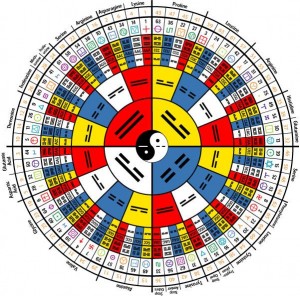
Oh, and I really like that codon wheel. Do you have a link to the picture on flickr, so I can add it as a favorite?
I agree with your opinion that concept of “Book of Life” is very much misleading, otherwise “Book of Life” is a book with enormous gaps or missing information all of which is implicit in the properties of environment in which gene operates and nature takes them for granted.
@Mr. Guniyn
I wasn’t referring so much to the integrity of the companies, but rather to the consumer need they are addressing: that people want to know their future. As for the issue of interpreting the results: the whole point was the danger of over-interpretation, hence the “entrail reading” phrase. This is, for example, why many countries mandate genetic testing for monogenic diseases only, where the results are unambiguous. Even so, they are presented to the paretns-to-be by a professional genetic counselor.
DTC genomics companies continue to operate legally and protect themselves from lawsuits by stating that their services are for entertainment and educational purposes only. But somehow I doubt that the main interest of their clientle lies in the gene that codes for the quality of their earwax.
Great piece, Iddo. I think one of the things to stress about companies like 23andMe is that they are trying to engage more people in learning about themselves and about biology. A lot of scientific knowledge is viewed by the public with a “meh, what does this have to do with ME?” attitude. So let’s tap into that. But present it in a way that is conducive to learning about the science behind it and avoid the black box quick fix infomercial that we are unfortunately starting to see (e.g. DNA-based dating services, screening babies for talents). That fact that this area is actually relevant to people’s daily lives does make caution prudent.
@shwu
Clients of personal genomics companies are people who can afford to pay $400-$1000 for SNP analyses and are interested enough in the power of genetics to reveal something about themselves, their futures and their pasts (some of these companies do genealogies as well). They are definitely not part of the “meh” crowd, at least not when it comes to genetics. Some of them may not be well-educated about genetics, but they do believe in its informational power, and are willing to shell out some money to receive this information. I believe that inasmuch DTC genomics are engaged in outreach activities, those are basically advertisements aimed at increasing their consumer base, I have nor problem with that.
The problem lies in this gap between enthusiasm for a scientific method,and ignorance of its mechanism and limitations. In a benign scenario, this gap may lead to over-interpretation of the information by the clients. In a more malign scenario, this can lead to corporate irresponsibility, bordering on or crossing the line into fraud and quackery.